Precision medicine for protein binding sites
Proteins interact with other molecules through their protein binding sites, which are functionally important regions on the protein surface. Each binding site usually binds one or a few specific molecules, the ligands (Figure 1). A ligand can be an ion, another protein, a nucleic acid, a small molecule, or a water molecule. By predicting the binding affinity of protein-ligand complexes with simulations, the efficiency of drug candidates can be assessed in silico, which is a topic that receives continued interest in drug discovery research.
Genetic variations such as sequence variants that occur in coding regions of genes may alter proteins’ amino acids and presumably affect protein function. It was found that disease-causing sequence variants are preferentially located at protein-protein interfaces rather than in noninterface regions of protein surfaces. Somatic sequence variants are known to cluster in oncogenic proteins, and in some oncoprotein-oncotarget pairs, significant enrichments of such mutations were found within protein-protein, protein-nucleic acid and protein-metal ion binding sites. As such, binding site sequence variants are of great interest to drug development chemists and clinicians who seek to predict an individual’s response to a drug, which is known as precision medicine.
Scoring the binding efficiency
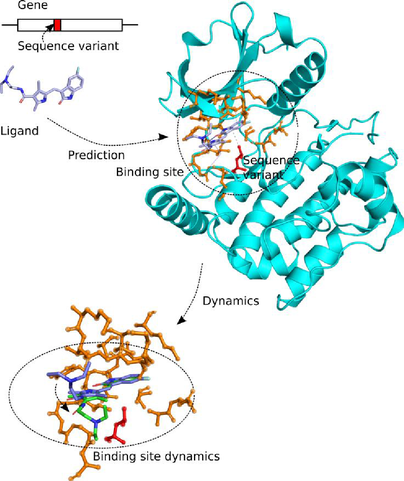
Precision medicine
Sequence variants can affect the binding affinity of small molecules to proteins and thus influence the individual's response to drug molecules. The ProBiS web server was developed by Konc et al. [1] which enables the detection of ligands for proteins, based on graph-theoretical matching criteria. The ProBiS algorithm was interfaced with molecular dynamics software, and extended to map sequence variants to proteins’ binding sites (GenProBiS). This will now be extended by systematic analysis of their influence on binding affinity and binding kinetics, using molecular dynamics simulations equipped with a Bayesian analysis methodology to fit trajectory data to the Smoluchowski diffusion model, using transition interface sampling and other techniques.
The developed approach will be validated on anti-cancer drugs, where the drugs sunitinib, imantinib, antibody cetuximab, and of small molecule drug warfarin will serve as test cases. Sunitinib is a cancer drug whose binding and selectivity towards protein targets is mediated by conserved waters and which has known issues in patients caused by its various modes of action due to the variability of human genome sequences: the presence and location of conserved structural water molecules in ligand binding sites will be registered and their importance for ligand binding will be assessed by determining their interactions with ligands.
The result will be novel computational solutions that will enable the prediction of pharmacodynamic and pharmacokinetic behaviour of active drug substances as a consequence of genetic variability.
IBiTech researchers currently active on the project
Funding sources
- Fund for Scientific Research – Flanders (FWO-Vlaanderen)
References
- J. Konc and D. Janežič, “ProBiS-ligands: A web server for prediction of ligands by examination of protein binding sites,” Nucleic Acids Res., vol. 42, no. W1, pp. 215–220, 2014